 Interest Statement: This work was funded in part by the Wm. Wrigley Jr. Company.
Interest Statement: This work was funded in part by the Wm. Wrigley Jr. Company.
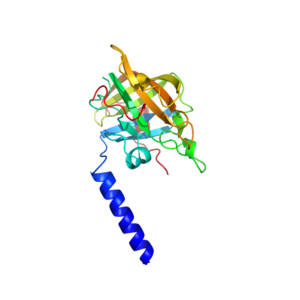
We recently determined the structure of the Streptococcus mutans sortase A protein in collaboration with the Campopiano group in the School of Chemistry at the University of Edinburgh. This work was part of the PhD project of one of Dom’s students who was working on natural product inhibition of bacterial pathogens and was sponsored by the Wm. Wrigley Jr. company. Our crystal structure was used to model the binding of trans-chalcone in the active site of the protein to help understand the mechanism of its inhibition of this protein. Sortases are found on the surface ofGram positive bacteria and attach certain proteins to the bacterial cell wall. These cell wall associated proteins are implicated in virulence and the formation of biofilms, and by inhibiting their attachment to the cell these proteins cannot function.
Trans-chalcone is a member of a huge family of flavonoid natural products that are found in liquorice and turmeric root to name just a few sources. In this paper we use biochemical assays and mass spectrometry to show that trans-chalcone irreversibly binds to Sortase A from the oral bacterial Streptococcus mutans. We modelled the interaction of this molecule with the crystal structure to show that larger chalcone molecules could be accommodated in the active site of this protein and that this could be used to select Sortase A specific inhibitors based on differences in the active-site cleft between the different classes of Sortase proteins.
This paper was reported by the Scottish Daily Mail:
The X-ray crystallographic data was all collected at Diamond Light Source as part of the University of Edinburgh BAG and was reported on the Diamond Light Source front page. Here’s the official University of Edinburgh press release.
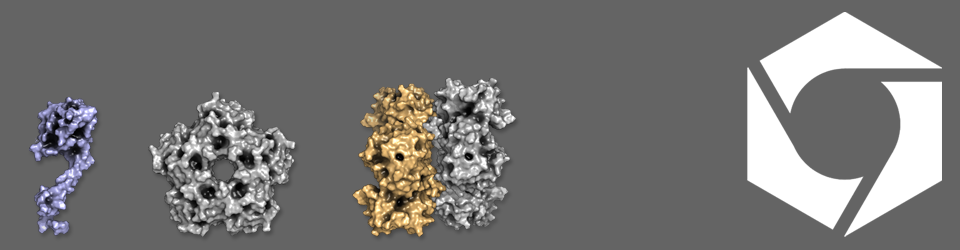

what’s Streptoccus mutans used for?
Not so much useful for biotech purposes. It’s an oral commensal and major cause of tooth decay.