From 21st-25th of May 2018, our PhD students Jasmine Bird and Andrea Giachino attended the STARS-sponsored training course “Demystifying Mats and Stats for Bioscientists” at Newcastle University, hosted by Prof Jarka Glassey. It was an invaluable development experience, packed full with state-of-the-art statistical techniques, experimental design, and private-sector golden standards.
Congratulations to Andrea for receiving the Best Presentation award (picture): his group’s presentation “Assessment of Bacterial Growth Parameters” showcased the implementation of advanced statistical methods to improve their research data.
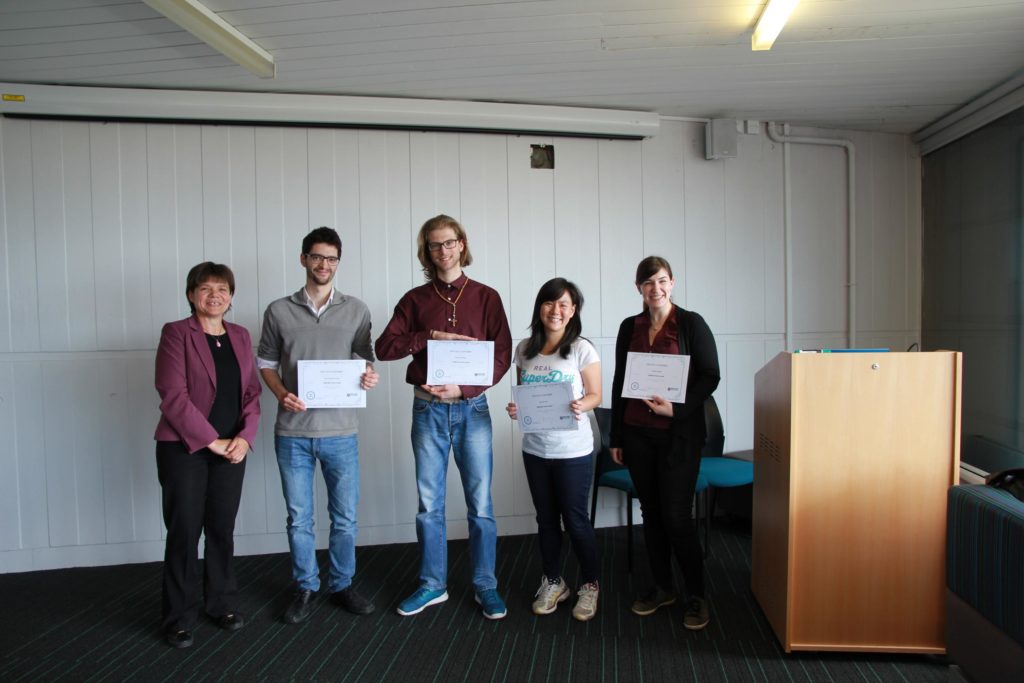

Best presentation award at “Demystifying Maths and Stats for Bioscientists” 2018. From left to right: Prof Jarka Glassey, Joan Cortada Garcia, Andrea Giachino, Gaik Sui Kee, and Martina Daute.
Andrea says:
I feel like I learnt so much from this course, and I will be looking at my data with different eyes from now on. It made a huge contribution to my skills as a researcher, improving my experimental design, data analysis, and critical thinking. What I liked the most was the amazing range of contributions from the private sector, which will benefit my long-term career goals. And of course, winning the final competition felt so rewarding!
Jasmine says:
I learnt a lot from this course, especially in terms of thinking about my data differently and more critically. This will benefit my data analysis in the future. The range of academic and industrial perspectives of modern problems in Biotechnology kept things interesting and gave me useful insight into how both areas tackle problems.